Chromatin | DNA Replication | Transcription | Genome Stability
It has not escaped our notice that the specific pairing we have postulated immediately suggests a possible copying mechanism for the genetic material.
Watson and Crick, Nature, 1953
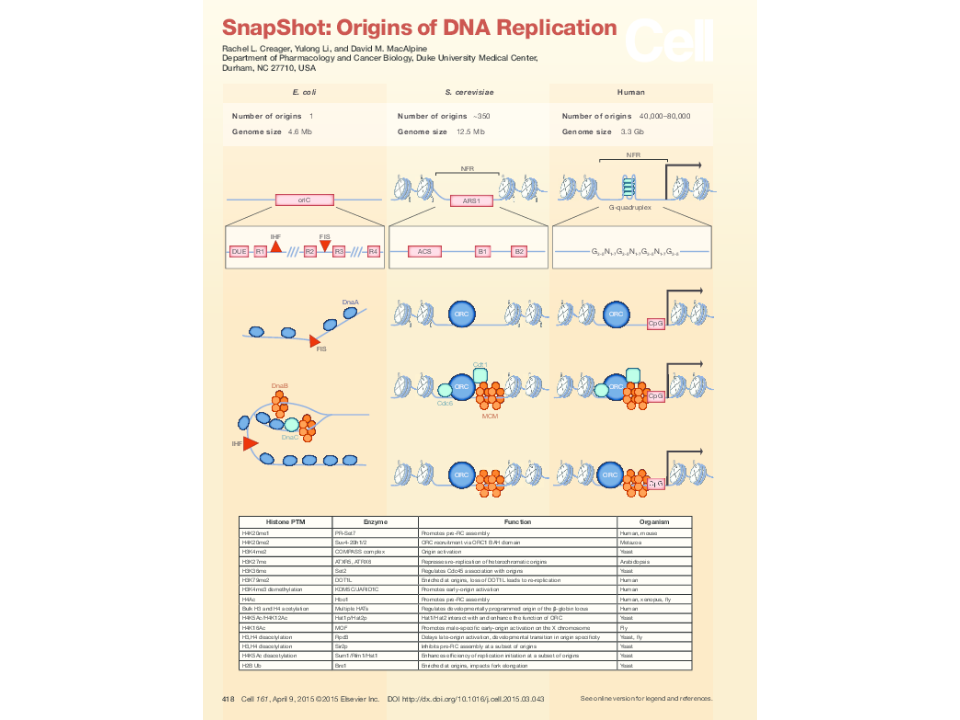
In the more than six decades since the discovery of the double helix, there has been tremendous progress in understanding the biochemical mechanisms responsible for the precise and accurate duplication of the genome. However, despite the remarkable conservation of proteins and protein functions required for DNA replication across both prokaryotic and eukaryotic systems, we know comparatively little about the functional elements that direct DNA replication in higher eukaryotes. My research program is focused on understanding how the start sites of DNA replication are selected and regulated in the context of the local chromatin environment to maintain genomic stability and to ensure the accurate inheritance of genetic and epigenetic information. Failure to accurately regulate DNA replication may result in under or over replication of the genome and lead to genomic instability -- a hallmark of tumorigenesis.
Research Projects

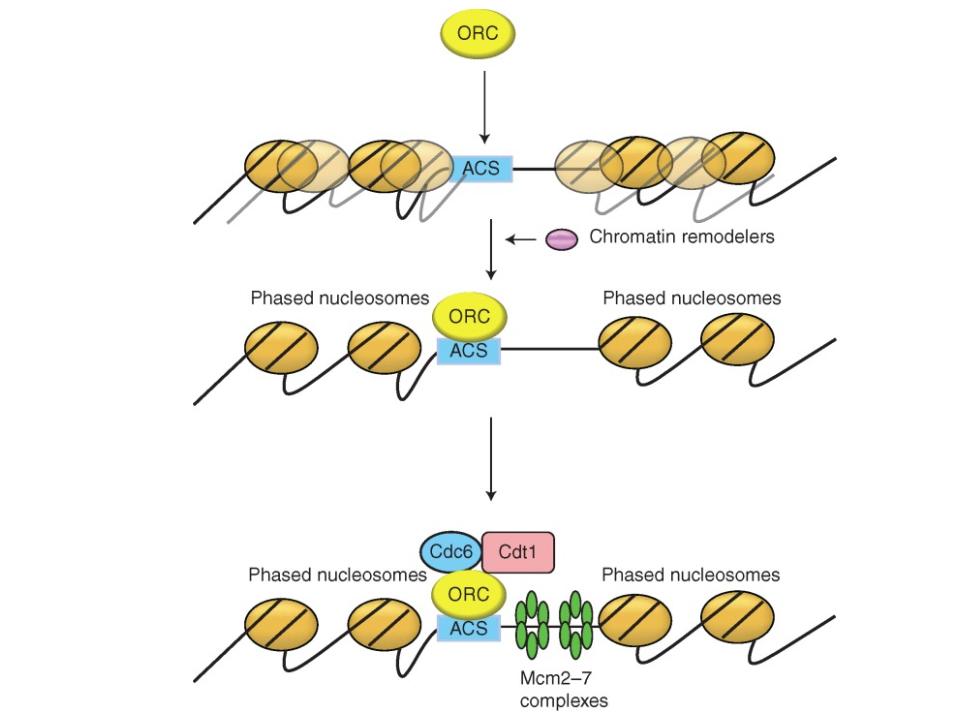
Chromatin remodeling at origins
Identification of chromatin remodeling activities that modulate origin function.
- Rachel Hoffman
- Heather MacAlpine
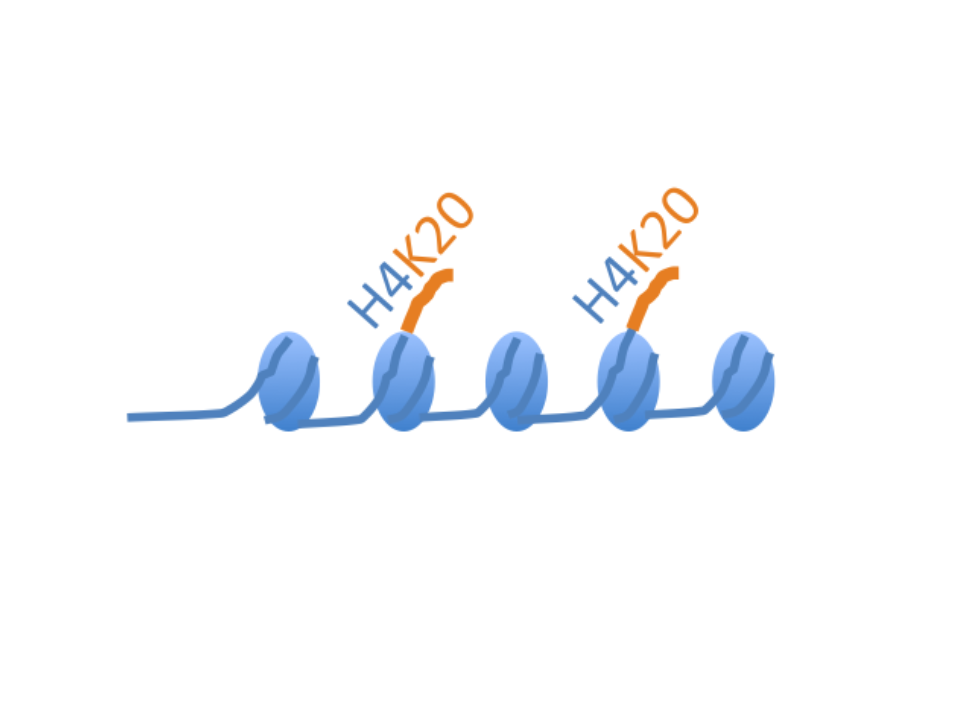
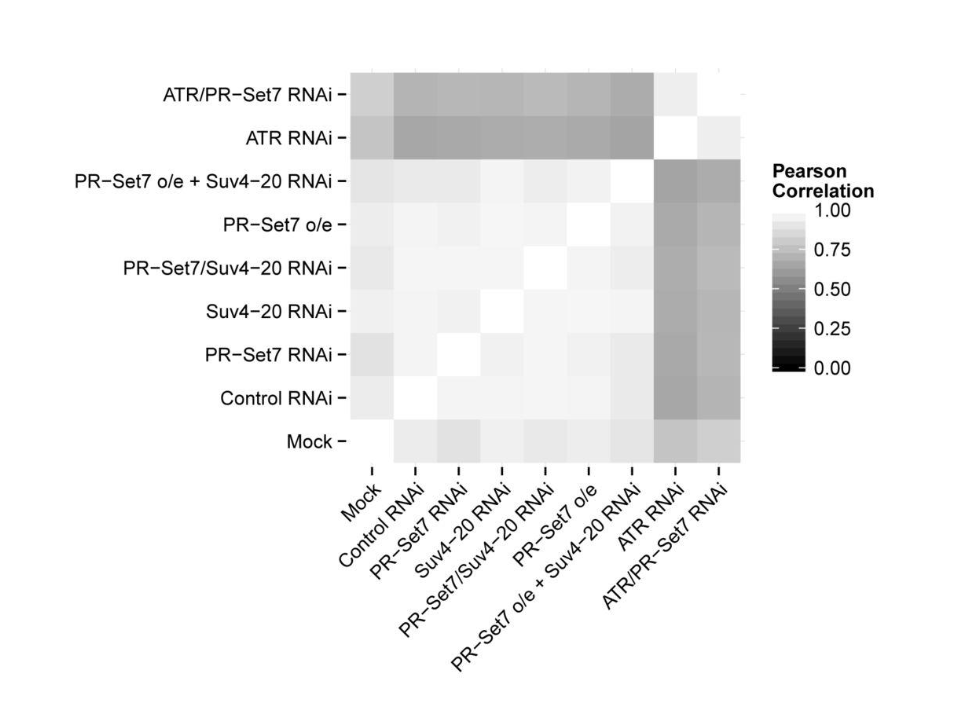
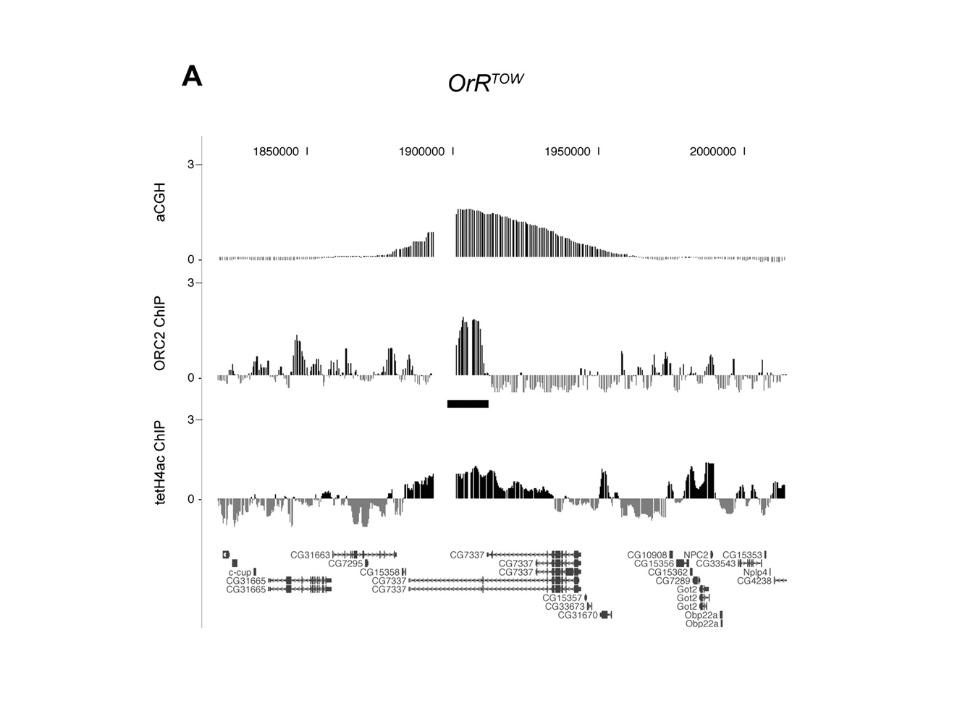
Chromatin structure and genome stability
Role of epigenetic writers and readers in maintaining the DNA replication program and genome stability.
- Yulong Li
- Duronio Lab (UNC)

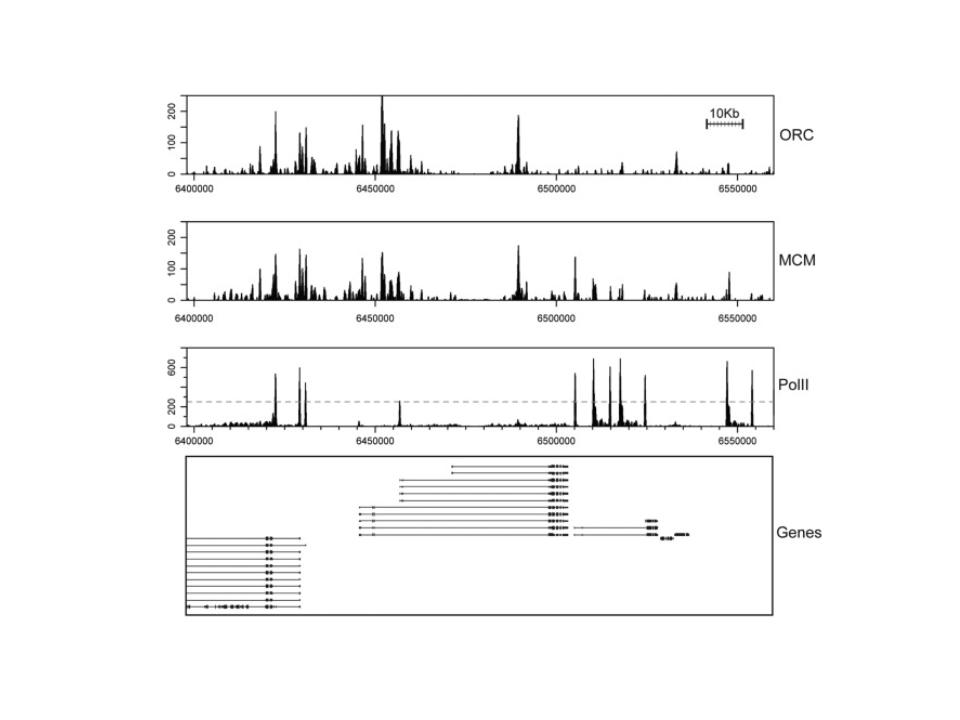
Predicting cell cycle gene expression from DNA occupancy
Development of experimental and computational approaches to predict cell cycle gene expression changes from chromatin occupancy data.
- Heather MacAlpine
- Vinay Tripuraneni
- Hartemink Lab (CS, Duke)

Chromatin structure of DNA breaks
Nucleotide resolution maps of chromatin structure following induction of site specific DNA breaks
- Vinay Tripuraneni
- Monica Gutierrez
- Sue Jinks-Robertson (MGM, Duke)
- Mike Resnick (NIEHS)
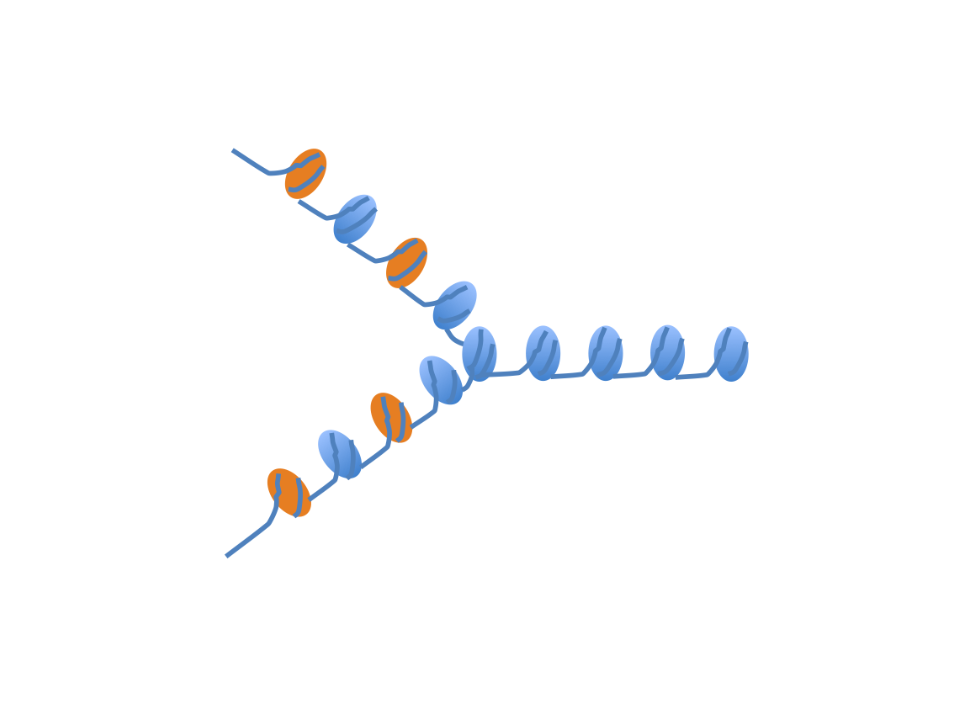
Chromatin assembly at the replication fork
Spatiotemporal re-assembly of chromatin behind the replication fork.
- Monica Gutierrez

Chromatin structure associated with biofilm formation
Developing chromatin occupancy based methods for mapping quantitative trait loci.
- Monica Gutierrez
- Cullen Roth (UPGG)
- Paul Magwene (Bio, Duke)
People
The MacAlpine laboratoy has diverse research interests from chromosome to computational biology. We have or have had graduate trainees from many of the graduate programs at Duke, including pharmacology, molecular cancer biology, cell and molecular bioliogy, genetics and genomics and computational biology.

My job is to keep the troops happy and well fed.
David MacAlpine Principal Investigator

My research focuses on the role of chromatin remodelers in establishing the DNA replication program. I'm also responsible for making sure that the lab runs efficiently.
Heather MacAlpine Laboratory Manager

I lounge around all day greeting people who walk by the lab. I will do anything for belly rubs and kibble.
Guinness MacAlpine Mascot

I am a UPGG graduate student and my research is focused on understanding how chromatin is assembled behind the replication fork to ensure the inheritance of epigenetic information.
Mónica Gutiérrez PhD Student, UPGG
Yulong Li is my name, bioinformatics is my game. I was a former MCB/CMB graduate student and now I am wrangling data and brushing up on my statistics.
Yulong Li Postdoc

I am a MSTP student also known as a mud phud. I tend to work the late shift while investigating chromatin structure following DNA damage.
Vinay Tripuraneni MD/PhD Student, MSTP
Where it all begins. I study the role of chromatin in regulating the initation of DNA replication.
Rachel Hoffman PhD Student, UPGG

Manners none. Ability to shred and destroy -- through the roof.
Islay MacAlpine Mascot in TrainingOur Publications
We use genomics and computational biology to address fundamental questions in chromosome biology.
Contact the Lab
Want to get more information? Contact us by using the form or call directly for more info.
-
Address
C333 LSRC, Duke University
-
Phone
+ 1 (919) 681 6077
-
Email
david.macalpine@duke.edu